MacVector
Native macOS Desktop Sequence Analysis Software
A complete molecular biology toolkit with over 35 years of continuous development
MacVector is the leading macOS desktop sequence analysis application, designed for molecular biologists to analyze, manipulate, assemble, and document DNA and protein sequences. Originally developed in the late 1980s by International Biotechnologies, Inc. (IBI), it has been under continuous development for over 35 years, making it one of the longest-running bioinformatics applications in existence.
MacVector provides a comprehensive all-in-one toolkit covering everything from primer design and cloning simulation, to restriction enzyme analysis and plasmid map drawing, to sequence assembly (Sanger, NGS, and long-read), protein analysis, and phylogenetic tree construction. The application is designed to be powerful yet intuitive, enabling bench scientists to perform complex bioinformatics tasks without command-line skills or bioinformatics expertise.
MacVector's key differentiator is its native macOS development — built as a true Cocoa application using Apple's frameworks, delivering a responsive, stable, Mac-native user experience. The software runs entirely offline (no cloud dependency), protecting proprietary sequences and supporting air-gapped laboratory environments. Since September 2024, MacVector includes the Assembler toolkit at no additional cost, providing six de novo assemblers and two reference mapping tools supporting all sequencing platforms.
Product Information
| Developer | MacVector, Inc. — Apex, North Carolina, USA (HQ) + Cambridge, UK (European office) |
| Platform | macOS only (native Cocoa application) |
| Latest Version | MacVector 18.8.2 (October 2025) |
| Licensing | Perpetual license (with optional maintenance renewal) or annual subscription |
| Free Version | MacVector Free (viewing, editing, basic file operations) |
Core Features
Sequence Editing & Visualization
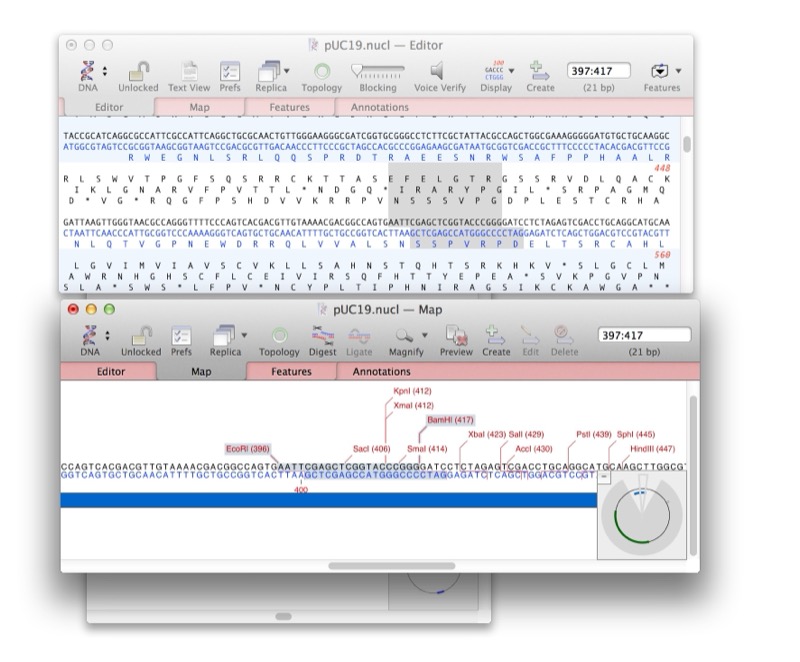
Rich Sequence Editor
Customizable feature display (colors, arrows, labels) with real-time feature annotation and editing
Interactive Map View
Circular and linear map views for plasmids and linear DNA, with publication-quality vector graphics export
Codon Usage Table Viewer
Codon usage analysis and optimization, with support for reverse translation and mRNA structure generation
Cloning & Construct Design
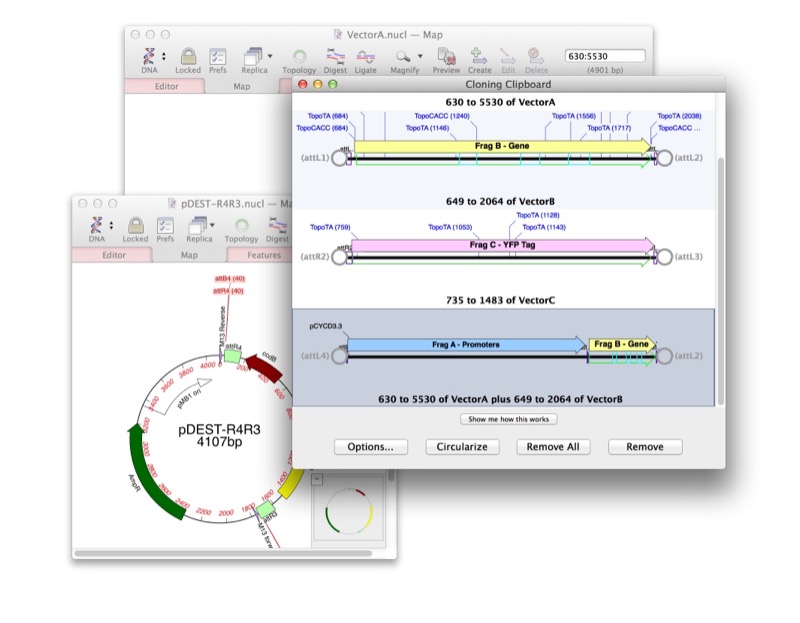
Click Cloning
Drag-and-drop interface for restriction enzyme cloning; Cloning Clipboard records all ligation events including enzymes used and dates
Gibson Assembly / LIC
Automated balanced primer generation for Gibson Assembly and Ligation Independent Cloning (LIC)
Gateway, TOPO & In-Fusion Cloning
Support for Invitrogen Gateway, TOPO TA, Zero Blunt, and Takara/Clontech In-Fusion technologies
Simulated Agarose Gel
Realistic restriction digest simulation with adjustable agarose concentration and electrophoresis time
Primer Design & Analysis
QuickTest Primer
Real-time interactive primer design with secondary structure preview, plus a primer database for cross-project storage and scanning
Comprehensive Primer Analysis
Tm calculation, GC content, self-complementarity analysis, primer pair optimization, and restriction site introduction into primers
Sequence Assembly (Assembler — Built-in Since September 2024)
De Novo Assemblers
Velvet (NGS), phrap (Sanger), and 4 other assemblers totaling 6, supporting Sanger, Illumina, 454, SOLiD, Helicos, Ion Torrent, PacBio, and Oxford Nanopore
Reference Assembly
Bowtie2 (NGS) and minimap2 (long-read), with interactive read alignment including SNP/INDEL detection and CRISPR validation
Assembly Project Management
Assembly Projects Manager, vector trimming (cross_match), contig editing, cDNA alignment, genome finishing, and gap closure tools

Sequence Search
BLAST & Entrez
Local and remote NCBI BLAST searches, with direct querying of NCBI databases
Restriction Enzyme & ORF Search
Complete restriction enzyme database search, ORF (Open Reading Frame) identification, motif search, and pattern matching
Multiple Sequence Alignment
Multiple Sequence Alignment
ClustalW, MUSCLE, and T-Coffee alignment algorithms, with MSA editor and consensus sequence generation
Dot Plot & Batch Scanning
Dot Plot optimized for bacterial genome comparison, and Align to Folder batch scanning for large datasets
Phylogenetic Analysis
Phylogenetic Tree Construction
Build phylogenetic trees from alignments with multiple tree-building methods, interactive tree visualization and editing, including Bootstrap analysis
Protein Analysis
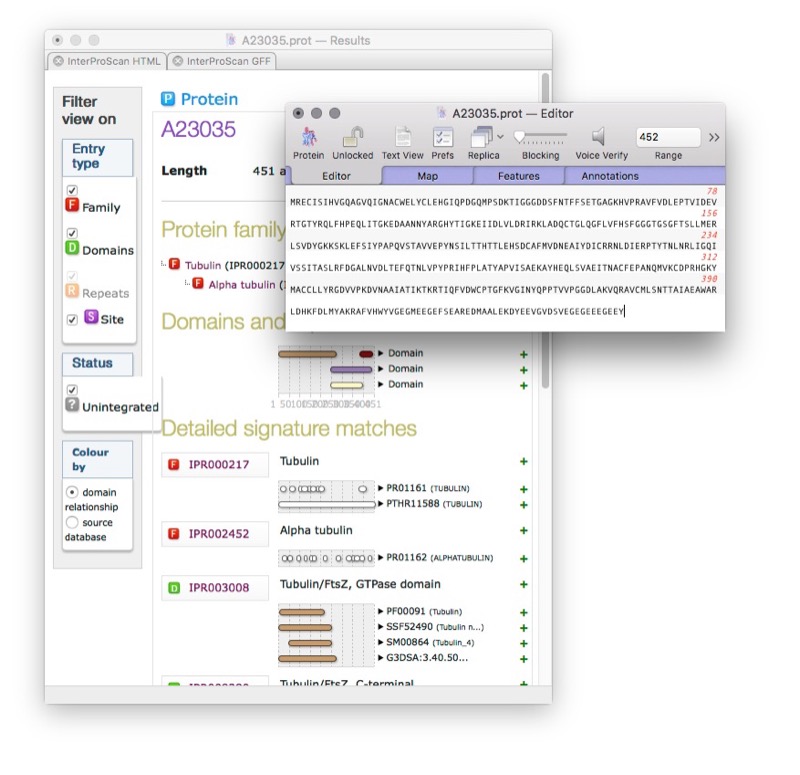
InterProScan Integration
Domain/family identification with graphical domain architecture display, annotating functional domains directly on proteins
Protein Structure & Property Analysis
Secondary structure prediction, hydrophobicity and antigenicity plots, signal peptide prediction, molecular weight and isoelectric point calculation
Technical Specifications
| Operating System | macOS only (native Cocoa application); supports current macOS versions |
| Architecture | 64-bit (since MacVector 14) |
| Runtime Environment | Fully offline — core features require no internet connection |
| Installation | Standard macOS .app bundle |
| License Activation | Serial number + activation code (standard) or hardware-locked (personal) |
Supported File Formats
| Import Formats | GenBank, EMBL, FASTA, FASTQ, SnapGene (.dna), SerialCloner (.xdna), SAM/BAM, BED, GFF/GFF3/GTF, PHYLIP, NEXUS, etc. |
| Export Formats | GenBank, EMBL, FASTA, PHYLIP, NEXUS, etc.; vector graphics (PDF, EPS) for publication-quality images |
| Sequencing Platforms | Sanger, Illumina, 454, SOLiD, Helicos, Ion Torrent, PacBio, Oxford Nanopore |
Company History
| Year | Event |
|---|---|
| Late 1980s | Developed by International Biotechnologies, Inc. (IBI) |
| Early 1990s | IBI acquired by Eastman Kodak Company; MacVector development continues |
| 1996 | Acquired by Oxford Molecular Group |
| 2001 | Oxford Molecular merged into Accelrys |
| 2007 | Dr. Kevin Kendall founded MacVector, Inc. for independent development and sales |
| 2025 | MacVector 18.8.2 released; over 35 years of continuous development |
Applications
Molecular Cloning
Restriction enzyme cloning, Gibson Assembly, Gateway/TOPO, In-Fusion — a complete suite of cloning planning and simulation tools.
Primer Design & Plasmid Mapping
PCR primer optimization, mutagenic primers, sequencing primer design; interactive circular/linear map creation with publication-quality output.
Sequence Confirmation & CRISPR Validation
Align Sanger sequencing reads to reference sequences for clone verification; use Align to Reference to detect CRISPR edits and INDELs.
NGS Data Assembly
De novo and reference assembly for short-read and long-read data, supporting all major sequencing platforms.
Protein Analysis & Phylogenetics
Domain identification, structure prediction, sequence annotation, as well as evolutionary analysis and tree construction.
Sequence Documentation
Lab-notebook-quality cloning experiment records; the Cloning Clipboard automatically logs dates, enzymes, and end modifications.
Key Differentiators
Native macOS Application
The only major sequence analysis software built as a true native macOS application — responsive, stable, Mac-optimized UI
Fully Offline Operation
No cloud dependency; core features require no internet connection, protecting proprietary sequences and supporting air-gapped lab environments
All-in-One + Built-in Assembler
A single application covering cloning, primer design, assembly, protein analysis, and phylogenetics. Assembler included free since September 2024
Perpetual License + 35+ Years of Development
Offers a perpetual license option (not subscription-only); licenses never expire. Designed for bench scientists with no bioinformatics expertise required
Long-Read Sequencing Support
PacBio and Oxford Nanopore assembly and reference mapping via minimap2
Target Users
Academic Research
Molecular biologists, geneticists, genomics researchers, graduate students and postdoctoral fellows, and academic research laboratories.
Industry & Government
Pharmaceutical and biotech R&D teams, clinical research laboratories, government research agencies (e.g., NIH, NCI, etc.).
Leading Global Institutions
Widely used at institutions such as Harvard, Stanford, UC Berkeley, NIH/NIAID, Massachusetts General Hospital, Max Delbruck Center, and more.
Related Products
Experience the Most Powerful Sequence Analysis Tool on Mac
Contact us to learn how MacVector can enhance your molecular biology research efficiency
Contact Us to Learn More