MetaCore / MetaDrug
Integrated manually curated knowledge database and pathway analysis software suite — Designed for multi-omics experimental data pathway analysis
MetaCore is Clarivate's integrated manually curated knowledge database and pathway analysis software suite, providing 100% manually curated and validated biological systems content, including molecular interactions defined by mechanism, directionality, and effect, enabling researchers to rapidly generate and validate hypotheses for novel biomarkers, drug targets, and mechanisms of action.
MetaCore's underlying knowledge base, MetaBase, is recognized as "the most comprehensive manually curated mammalian biology database on the market today" and is the only database that simultaneously provides directionality, mechanism, and effect attributes for all interactions. The companion product MetaDrug provides a systems pharmacology solution for predicting metabolism, toxicity, and biological effects of small molecule compounds.

MetaCore — Analyze molecular pathways and accelerate discovery research, covering 4M+ manually curated molecular interactions and 1,500+ pathway maps
MetaBase Knowledge Base Coverage
Molecular Interactions
100% manually curated, each interaction with directionality, mechanism, and effect attributes — the only one on the market
Canonical Pathway Maps
Signaling and metabolic pathway maps, with 1,100+ manually curated and validated networks
Gene-Disease Associations
Gene-disease association analysis covering 10,000+ diseases
Compounds
Compounds with targets and bioactivity, along with approximately 8,900 drug records
Metabolic Reactions
Including 5,300+ endogenous metabolites, supporting metabolomics analysis
Publication References
Continuously curated and updated by PhD and MD research professionals (25+ years of experience)
Core Features
Pathway Enrichment Analysis
Map experimental data to 1,500+ curated canonical pathway maps using hypergeometric distribution for statistical enrichment. Supports ontology enrichment across MetaCore pathway maps, GO cellular processes, disease biomarker networks, drug target networks, and toxicity networks
Network Analysis
10+ network generation algorithms with tissue, functional process, and species-specific filters. Direct interaction and shortest path algorithms, network scoring using G-scores and P-values, with 6 interactome tools
Key Pathway Advisor (KPA)
Drag-and-drop web application requiring no bioinformatics expertise. Predicts key protein activity changes (root cause analysis) using the SPIA algorithm, delivering results in 2 steps and under 1 minute
Multi-Omics Data Integration
Supports transcriptomics, proteomics, metabolomics, genomics, epigenomics, and functional genomics. Universal parser supports Affymetrix, Agilent, Illumina, and SNP chip formats
Visualization & Custom Editing
Interactive canonical pathway maps with Pathway Map Creator for drawing/editing custom pathway maps. All features support custom network editing, with dynamic graphs for time-series data
100% Manual Curation
All interactions manually curated and validated by PhD and MD-level experts. No NLP or automated extraction, with industry-leading ontology ensuring annotation consistency
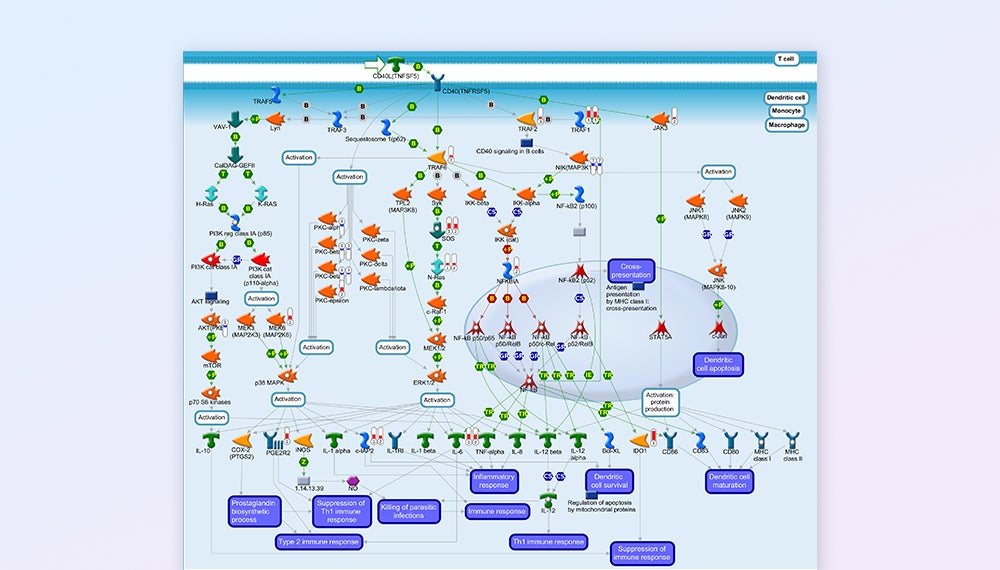
MetaCore Pathway Map Visualization — CD40 signaling pathway covering immune response, dendritic cell, and macrophage signaling networks
MetaDrug Companion Product
MetaDrug is Clarivate's systems pharmacology solution for predicting metabolism, toxicity, and biological effects of small molecule compounds. It won the Best New Product Award at the Molecular Medicine Conference in 2007.
Metabolite Prediction
10,000+ xenobiotic reactions, 80 key metabolic reaction rules
ADME/Tox Prediction
QSAR models for 40+ major drug-metabolizing enzymes
Activity Prediction
Predict activity of compounds and their metabolites
Off-Target Effects
System-level chemoinformatics tools combining compound-target interactions
Indication Prediction
Predict potential therapeutic indications for novel small molecules
Pathway Integration
Integrate prediction results with human cell signaling and metabolic pathways
Drug Development Lifecycle Coverage
| Stage | MetaCore Capabilities |
|---|---|
| Target Exploration | Disease-gene association analysis (182K+), network-based target identification, interactome analysis |
| Target Validation | Pathway enrichment to confirm biological relevance, causal reasoning analysis |
| Lead Identification | Compound-target interaction mapping (750K+ compounds), off-target effect prediction via MetaDrug |
| Preclinical / Safety | ADME/Tox prediction, toxicogenomics pathway analysis, hepatotoxicant assessment |
| Biomarker Development | Diagnostic and prognostic biomarker identification throughout the development cycle |
| Drug Repositioning | Network-based identification of new indications for existing drugs (e.g., COVID-19 research identified 15 drug targets and 46 existing drugs) |
Therapeutic Expertise Areas
Oncology
Breast cancer GWAS, glioblastoma, ovarian cancer, liver cancer, chronic lymphocytic leukemia
Immunology
Immune response pathway enrichment, vaccine research
Infectious Diseases
COVID-19 drug repurposing research
Rare Diseases
MetaMiner Cystic Fibrosis — the first disease-specific software platform
Toxicology
Hepatotoxicant research, drug-induced toxicity assessment
CNS
Disease ontology and curated neurodegenerative disease networks
Core Competitive Advantages
The Only Database with All Three Interaction Attributes
Directionality, mechanism, effect — the only database on the market with all three attributes simultaneously
100% Manual Curation
No NLP or automated extraction; all interactions validated by PhD-level experts, ensuring the highest data quality
Deep ADME/Tox Integration
Metabolite prediction, safety assessment, and indication prediction via the MetaDrug companion product
Cortellis Ecosystem Integration
End-to-end drug discovery with seamless integration with CDDI, CCI, and other Cortellis modules, single sign-on access
Industries & Users
| Industry | Primary Use Cases |
|---|---|
| Pharmaceutical Companies | Target screening/validation, biomarker identification, mechanism of action research, drug repositioning |
| Biotech Companies | Drug discovery pathway analysis, lead optimization, translational research |
| Government / Regulatory | U.S. FDA authorized to use MetaCore for reviewing genomics data in drug applications |
| Academic / Research Institutions | Systems biology research, multi-omics analysis (NIH, MIT, Yale, Stanford, Harvard, etc.) |
| Contract Research Organizations | Genomics services for biotech/pharma clients |
| Disease Foundations | Disease-specific platform development (e.g., Cystic Fibrosis Foundation) |
Supported Input Data Types
| Data Type | Specific Formats |
|---|---|
| Transcriptomics | Affymetrix, Agilent, Illumina (microarray); RNA-seq / NGS, SAGE, MPSS (sequencing) |
| Proteomics | Protein expression data, Co-IP pull-out |
| Metabolomics | Metabolite lists, supporting enrichment and pathway analysis |
| Genomics | SNPs, CGH chips, genetic variants |
| Epigenomics | ChIP-seq data |
| Functional Genomics | RNAi screens, siRNA, microRNA |
Related Products
Learn About MetaCore / MetaDrug
Power your multi-omics research and drug discovery pathway analysis with a 100% manually curated knowledge base
Contact Us