BIOVIA Discovery Studio
Life Science Molecular Simulation Platform — A complete computational drug discovery solution from target identification to preclinical development
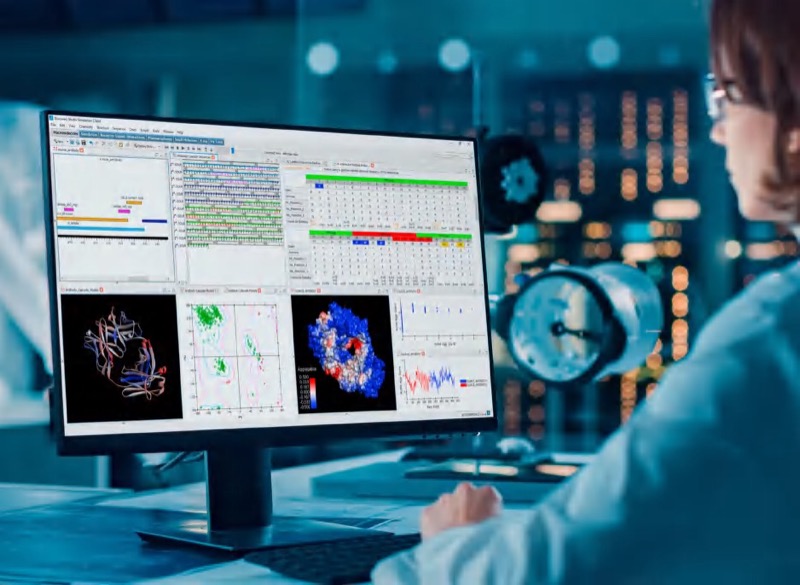

BIOVIA Discovery Studio is the most comprehensive 3D modeling and simulation application for drug discovery research in the life sciences. Built on the BIOVIA Pipeline Pilot infrastructure, Discovery Studio integrates over 30 years of industry research and world-leading in silico technologies within a single interface, providing a comprehensive solution that supports early-stage R&D for both biotherapeutics and small molecule drugs.
The platform integrates methods developed and validated by academic leaders at Harvard University, MIT, and UCSF, and works with proven advanced simulation programs such as CHARMm, NAMD, MODELER, ZDOCK, GOLD, and Catalyst. From early target identification through lead discovery and optimization to preclinical formulation development, Discovery Studio addresses all computational needs throughout the drug discovery process.
| Product Category | Computational Chemistry / Molecular Modeling & Simulation |
| Developer | Dassault Systèmes, BIOVIA brand |
| Platform Type | Windows desktop application + server-side computation; 3DEXPERIENCE platform |
| Built On | BIOVIA Pipeline Pilot |
| History | Originally Accelrys (founded 1981), acquired by Dassault Systèmes in 2014 |
Core Features
Protein Modeling & Engineering
MODELER Homology Modeling
Uses the market-leading MODELER homology modeling algorithm to rapidly predict high-quality 3D protein models from amino acid sequences; DS 2026 includes MODELER v10.7
Sequence Analysis & Property Calculation
BLAST and PSI-BLAST sequence searches, feature/motif prediction, post-translational modification (PTM) site prediction, and protein ionization and isoelectric point calculations
Loop Region & Side Chain Optimization
Uses CHARMm's LOOPER algorithm for systematic loop conformation sampling and optimization; ChiRotor for amino acid side chain position optimization
ZDOCK Protein-Protein Docking
Uses ZDOCK for protein-protein docking studies with result refinement; supports combinatorial amino acid mutation to assess stability and binding affinity
Biotherapeutics & Antibody Modeling
Antibody Structure Prediction
Supports IMGT, Kabat, Chothia, and Honegger numbering schemes; generates high-quality full-length antibody, Fab, or Fv 3D structures from light and heavy chain sequences (DS 2026 includes 8,083 PDB antibody templates)
Developability Assessment
Computes solubility, viscosity, and developability index (DI); uses Surface Charge Method (SCM) for viscosity estimation and AggMap aggregation propensity scoring; predicts preferential interactions for six common excipients
Humanization & Affinity Maturation
Antibody humanization design to reduce immunogenicity; paratope residue prediction (DS 2025 new feature); antibody-antigen complex creation and affinity maturation studies (DS 2025 new feature)
Simulations
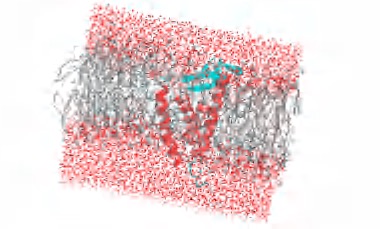
CHARMm & NAMD Molecular Dynamics
Performs energy minimization and MD simulations on GPU and CPU; supports CGenFF and charmm36 force fields; NAMD GPU-resident mode (DS 2025); OpenMM GPU acceleration (DS 2026)
MSLD Free Energy Calculations
Multi-Site Lambda Dynamics computes relative binding free energies for an entire combinatorial library in a single GPU simulation, 20x more efficient than traditional FEP; ideal for large-scale lead optimization
FEP & QM/MM
Free energy perturbation simulations to compute relative free energies for multiple ligand pairs; OpenFE framework integration (DS 2026); combined DMol3 and CHARMm for QM/MM studies

Structure-Based & Fragment-Based Design
Multiple Molecular Docking Methods
CDOCKER (CHARMm), LibDock (hotspots), Ludi, MCSS, and pharmacophore methods; interactive docking with CCDC GOLD (DS 2026 includes GOLD v2025.1)
Lead Optimization
In-situ optimization using classical medicinal chemistry reactions with commercial reagents; scaffold hopping or R-group substitution for new molecular designs; MM-PBSA/MM-GBSA binding energy calculations
Pharmacophore-Based & Ligand-Based Design
Catalyst Pharmacophore Engine
Market-leading automatic 3D pharmacophore generation (active ligands, receptor sites, receptor-ligand complexes); paired with PharmaDB library (41,000+ entries) for off-target and drug repurposing exploration
Combinatorial Library & Screening Optimization
Reaction-based enumeration or Markush combinatorial libraries; library selection optimization using Pareto optimization, clustering, diversity, and similarity analysis
ADMET / TOPKAT Prediction
ADMET Property Prediction
Evaluates blood-brain barrier penetration, human intestinal absorption, aqueous solubility, hepatotoxicity, CYP2D6, and other pharmacokinetic properties; supports hundreds of physicochemical and quantum mechanical descriptors
TOPKAT Toxicity Assessment
Ames mutagenicity, rodent carcinogenicity, skin irritancy and sensitization, ocular irritancy, aerobic biodegradability; used by Labcorp for agrochemical regulatory toxicology assessment
Integrated Engines
| Engine | Function |
|---|---|
| CHARMm | Force field-based simulation (CPU + GPU versions) |
| NAMD | Force field-based simulation (CPU + GPU versions) |
| MODELER | Protein homology modeling (v10.7 in DS 2026) |
| BLAST+ | Sequence search |
| GOLD | Protein-ligand docking (CCDC, v2025.1 in DS 2026) |
| ZDOCK | Protein-protein docking |
| Catalyst | Pharmacophore modeling |
| AggMap / SCM | Protein aggregation and viscosity prediction |
| CDOCKER | CHARMm-based flexible molecular docking |
| LibDock | Hotspot-based docking |
| DMol3 | Quantum mechanics calculations (QM/MM) |
| TOPKAT | QSAR predictive toxicology |
Technical Specifications
| Platform | Windows desktop client + server computation |
| Server OS | Windows Server; Linux support removed in DS 2026 |
| GPU Support | NVIDIA GPU for CHARMm, NAMD, MSLD, FEP; OpenMM acceleration (DS 2026) |
| Infrastructure | Built on BIOVIA Pipeline Pilot (requires Pipeline Pilot installation) |
| Force Fields | CGenFF, charmm36, CHARMm |
| File Formats | PDB, mmCIF (DS 2026 full multi-character chain name support), SDF, MOL2 |
| Databases | PharmaDB (41,000+ entries, scPDB 2024), curated PDB antibody templates (8,083 structures) |
| Integration | 3DEXPERIENCE platform, BIOVIA Pipeline Pilot, GOLD (CCDC) |
| Free Tool | BIOVIA Discovery Studio Visualizer available for free download |
Application Areas
Pharmaceutical Companies
Small molecule drug discovery, lead optimization, virtual screening, ADMET prediction
Biotechnology Companies
Antibody design, biologic developability assessment, protein engineering
Academic / Research Institutions
Structural biology, computational chemistry, drug discovery training
Agrochemicals / Government Agencies
TOPKAT regulatory toxicology prediction (used by Labcorp); COVID-19 drug repurposing research
Drug Development Lifecycle Coverage
| Stage | Key Tools & Methods |
|---|---|
| Target Identification | Homology modeling (MODELER), protein-protein docking (ZDOCK), sequence analysis |
| Lead Identification | Virtual screening, pharmacophore screening (Catalyst), fragment-based design |
| Lead Optimization | MSLD/FEP binding free energy, scaffold hopping, in-situ R-group substitution, QSAR |
| Preclinical | ADMET prediction, TOPKAT toxicity, mutagenicity, carcinogenicity assessment |
| Biologics Development | Antibody modeling, humanization, developability (DI, viscosity, aggregation), formulation |
| Formulation Development | Excipient prediction, protein stability assessment |
Key Competitive Advantages
Most Comprehensive All-in-One Platform
Biologics + small molecules + ADMET + simulation integrated in a single interface
MSLD High-Efficiency Free Energy Calculations
20x more efficient combinatorial library free energy calculations compared to traditional FEP
Market-Leading Core Engines
Catalyst pharmacophore engine, MODELER homology modeling, TOPKAT toxicology — each an industry benchmark in its field
Enterprise Integration & Automation
Pipeline Pilot foundation enables cross-BIOVIA product workflow automation; 3DEXPERIENCE platform for end-to-end R&D lifecycle management
30+ Years of Scientific Validation
Tens of thousands of peer-reviewed references; methods validated by top academic institutions including Harvard, MIT, and UCSF
Recent Updates
| Date | Update |
|---|---|
| November 2025 | Discovery Studio 2026 — OpenMM GPU acceleration, OpenFE free energy framework, enhanced pocket detection and allosteric motion analysis, mmCIF multi-character chain support, MODELER 10.7, GOLD 2025.1; Linux client removed |
| November 2024 | Discovery Studio 2025 — Antibody paratope prediction, PharmaDB 41,000+ entries (scPDB 2024), NAMD GPU-resident mode, full mmCIF support, CHARMm extended to 1 million atoms |
Related Products
Learn About Discovery Studio
Discover how molecular simulation can accelerate your drug discovery process
Contact Us